Fine-tune generated molecular poses with a force field
Published:
Some molecular pose generation methods benefit from an energy relaxation post-processing step.
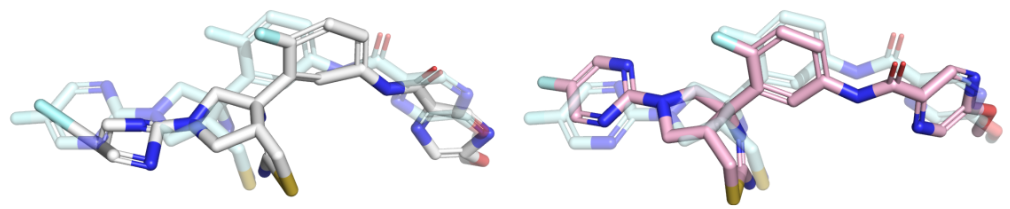
Here is a quick way to do this using OpenMM via a short script I prepared:
Get script
git clone https://github.com/maabuu/posebusters_em.git
cd posebusters_em
Setup environment
conda create -n posebusters_em openff-forcefields openff-interchange openff-toolkit "openmm>=8" "openmmforcefields>=0.11" pdbfixer "rdkit>=2022" -c conda-forge
conda activate posebusters_em
Run minimization
python energy_minimization.py test_cases/7MYU_ZR7/7MYU_ZR7_protein.pdb test_cases/7MYU_ZR7/7MYU_ZR7_prediction.sdf 7MYU_ZR7_prediction_minimized.sdf -t cache_dir
This example takes ~2 minutes to run on a desktop computer. Depending on the system’s complexity, runs may take much longer.
The script is available on my GitHub.
